"Because of its great advantages of the high sensitivity and specificity, real-time PCR becomes the 'gold standard' in the detection of SARS-CoV-2. The capacity of producing millions of copies of a specific target from a few initial copies also brings the researchers a headache problem, contamination."

The contamination issues can result in false positive results,moreover the contamination is very hard to be reduced or removed once it has occurred.
Therefore, to prevent the occurrence of contamination is very important for a PCR Lab. A proper laboratory setup and workflow, together with Good Laboratory Practice will greatly help to reduce the likelihood of contamination. In this article, we summarized some tips in setup a PCR lab and some basic practices and procedures for avoiding contamination.
PCR Laboratory Set-Up
1. Physical separation of work areas
A laboratory performing PCR analyses on diagnostic samples should be divided into at least three physically separate rooms (Figure 1):
- Reagent preparation area
- Sample preparation area
- Amplification and product detection area
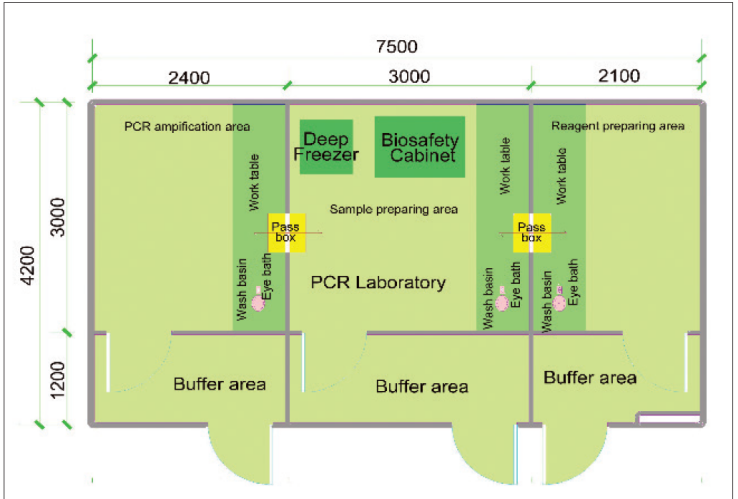
Figure 1
The reagent preparation area is for reagent aliquoting and mastermix preparation. This room should be the cleanest of all spaces used for the preparation of PCR experiments and should ideally be in a designated laminar flow cabinet equipped with a UV light. PCR reagents should be kept in a refrigerator (as per manufacturer recommendations) in the same designated area, ideally next to the laminar flow cabinet. The Master-mix should be prepared and aliquoted in the reaction vessel and extracted nucleic acid and amplified PCR products must not be handled in this area.
Sample preparation area is for nucleic acid extraction and DNA template and controls addition. When adding the samples, it should ideally be in a designated biosafety cabinet equipped with a UV light. To avoid contamination of the extracted nucleic acid samples that are being analysed, it is recommended to firstly add sample and negative control, then lastly add positive controls or standards. Samples should be stored in designated fridges or freezers in the same area and PCR reagents or amplified products must not be pipetted in this area.
Amplification and product detection area is for amplification and handling of amplified product, and product analysis (not for real-time PCR). It usually contains thermocyclers and real-time platforms. Ideally a laminar flow cabinet would be used for any steps that require manipulating open tubes after PCR amplification.
2. Air pressure and air conditioning
- Reagent preparation area (using positive air pressure to prevent the introduction of contamination)
- Sample preparation are (using negative air pressure to keep template nucleic acids in the room)
- Amplification and product detection area (using negative pressure to keep amplified nucleic acids in the room)
- The air handlers for the three areas need to be connected to separate air ducts, and each must lead to a separate location for exhaust. Do NOT use central air-conditioning system and ductwork for the three areas.
3. Unidirectional workflow
When a PCR procedure starts, the direction will be Reagent Preparing Area → Sample Preparing Area → PCR Amplification Area. NEVER return previously processed tubes to preceding work area or room. Operators who have worked in post-PCR should not go back and work in pre-PCR. Also, the consumables and PPE (lab coats, gloves, goggles, etc.) that have been introduced into the post-PCR room should never be placed back in the pre-PCR room without thorough decontamination. Waste materials that contain PCR amplicons should not be allowed to accumulate in an area close to the personnel involved with template isolation and purification.
4. Dedicated equipment and consumables
Besides of the separated spaces, all the supplies and equipment such as centrifuges, storage freezers/refrigerators, pipettors, reagents, pipettor tips, racks and so forth should be dedicated to each area and should not be interchanged between areas. Also, the lab coats for operator should be dedicated for three areas separately.
PCR Laboratory Operation
1. Surface and equipment decontamination
Contamination of work surfaces and equipment will be reduced by using 70% ethanol to clean them before and after PCR. Regularly use a freshly made 10–15% sodium hypochlorite to clean the surface and wait for 10 to 15 minutes before wiping down with de-ionized water. Alternatively, commercially available products that are validated as DNA-destroying surface decontaminants can be used if sodium hypochlorite is not suitable for decontaminating the metallic parts of equipment. If using 70% ethanol instead of sodium hypochlorite, irradiation with UV light will be needed to complete the decontamination.
2. UV light
Before use, the laminar flow cabinet or biosafety cabinet should be decontaminated using UV light for at least half an hour and cleaned with 70% ethanol. The airflow and HEPA filtration in all hoods should be monitored and certified as per manufacturers’ recommendations. Regularly expose the working areas and room to UV light overnight.
3. Using aerosol barrier tips
Aerosols can lead to cross-contamination from sample-to-sample. Aerosol barrier pipette tips could prevent the reintroduction of small amounts of a contaminating aerosolized sample into the next sample that is pipetted. In addition, correct pipetting technique also helps to reduce contamination. Incorrect pipetting may result in splashing when dispensing liquids and the creation of aerosols.
4. Carefully open and close sample tubes
All sample tubes and reaction plates should be opened and closed very carefully to make sure the liquid doesn’t splash out. Spinning tubes/plates before opening can prevent aerosols when opening.
5. Setup controls on every PCR experiment
Include well-characterized, confirmed positive and negative controls, along with a no-template control in all reactions. The positive control should not be very high concentration due to it poses a contamination risk. A no-template control (NTC) is used to check for the absence of contamination in the reagents, consumables, and environment.
6. Uracil-DNA-glycosylase (UDG) / uracil-N-glycosylase (UNG)
The use of UDG/UNG and dUTP substituted for dTTP could help to remove carryover amplification contamination from the previous PCR reactions. The UDG/UNG method works best with T-rich amplification products and is not as effective with GC-rich amplification products. UDG/UNG is not effective for sources of DNA contamination, other than dUTP-containing amplification products from previous qPCR experiments.
Conclusion
This article listed several common approaches to reduce or avoid contamination in PCR laboratory for your reference. You may refer to the above tips and combine with your local guidance and good laboratory practice to prevent the occur of contamination.
Reference
- “Establishment of PCR laboratory in developing countries 2nd version." N.p, n.d. 2016
- Mifflin TE. Setting up a PCR laboratory. CSH Protoc. 2007 Jul 1; 2007: pdb. top14. doi: 10.1101/pdb. top 14. PMID: 21357132.
- Ausubel FM. The polymerase chain reaction. In: Ausubel F, Brent R, Moore D, Seidman JG, Smith J, Struhl K, editors. Current Protocols in Molecular Biology, John Wiley & Sons, Inc., U.S.A., Vol. 2, Chapter 15, 1999; 15.01
- Aslanzadeh J (2004) Preventing PCR amplification carryover contamination in a clinical laboratory. Ann Clin Lab Sci 34(4):389–96

